A crustacean annotated transcriptome (CAT) database
Background
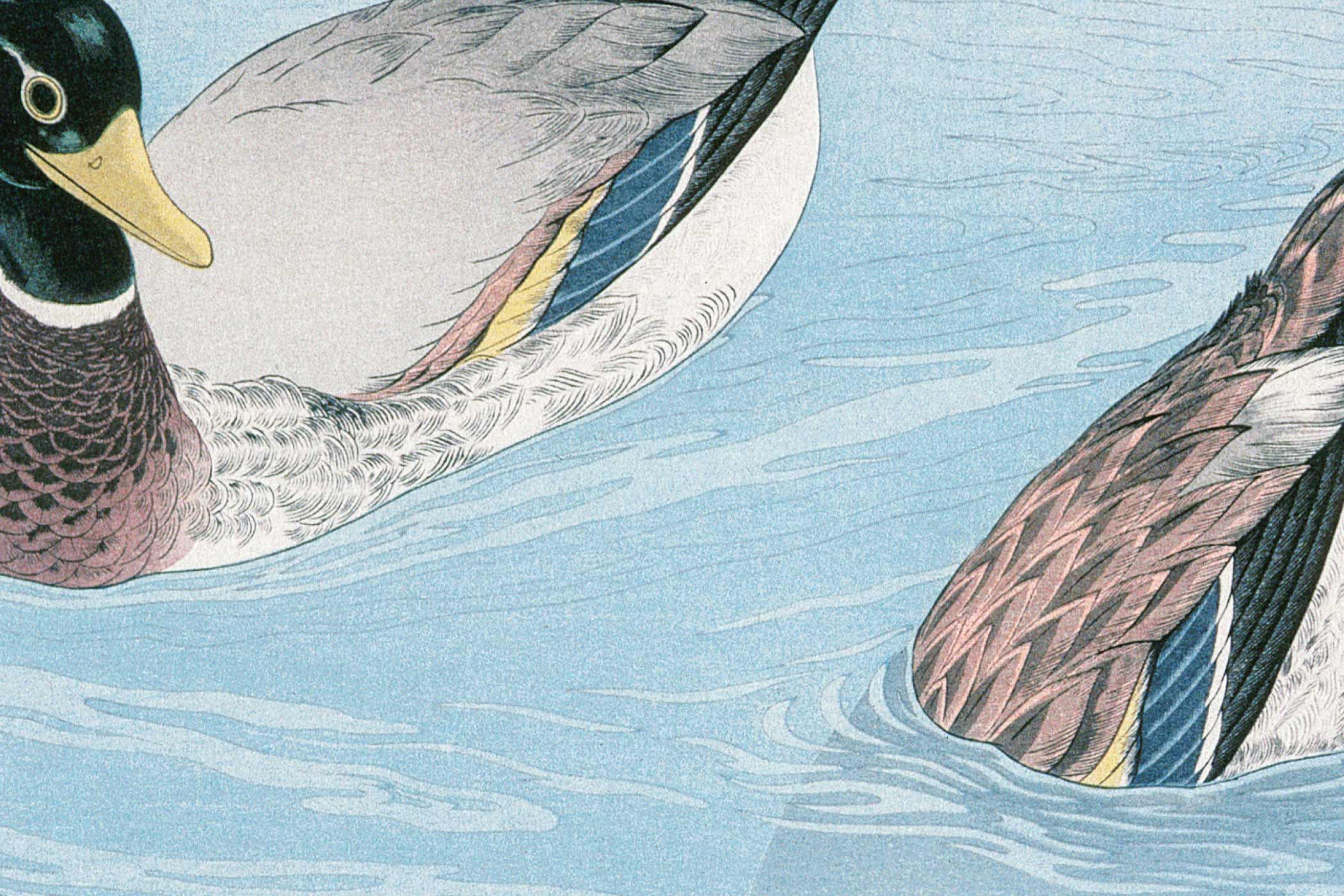
Decapods are an order of crustaceans which includes shrimps, crabs, lobsters and crayfish. They occur worldwide and are of great scientific interest as well as being of ecological and economic importance in fisheries and aquaculture. However, our knowledge of their biology mainly comes from the group which is most closely related to crustaceans – insects. Here we produce a de novo transcriptome database, crustacean annotated transcriptome (CAT) database, spanning multiple tissues and the life stages of seven crustaceans.
Description
A total of 71 transcriptome assemblies from six decapod species and a stomatopod species, including the coral shrimp Stenopus hispidus, the cherry shrimp Neocaridina davidi, the redclaw crayfish Cherax quadricarinatus, the spiny lobster Panulirus ornatus, the red king crab Paralithodes camtschaticus, the coconut crab Birgus latro, and the zebra mantis shrimp Lysiosquillina maculata, were generated. Differential gene expression analyses within species were generated as a reference and included in a graphical user interface database at http://cat.sls.cuhk.edu.hk/. Users can carry out gene name searches and also access gene sequences based on a sequence query using the BLAST search function.
Conclusions
The data generated and deposited in this database offers a valuable resource for the further study of these crustaceans, as well as being of use in aquaculture development.
Reference
- Nong WY, Chai ZY, Jiang X, Qin J, Ma KY, Chan KM, Chan TF, Chow BK, Kwan HS, Wong CK, Qiu JW, Hui JHL*, Chu KH. (2020). A crustacean annotated transcriptome (CAT) database. BMC Genomics, 21, 32. (Link)